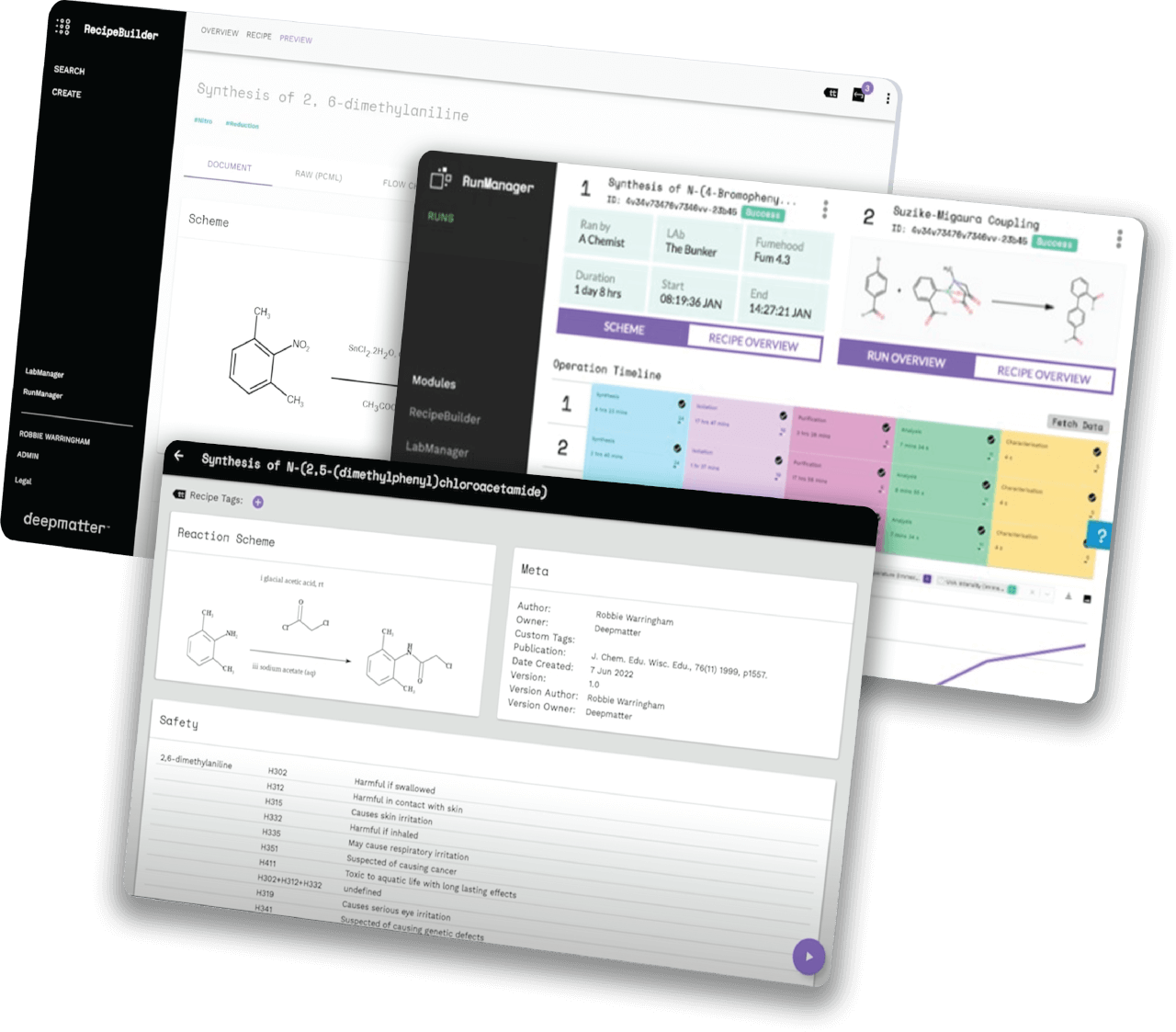
Helping companies worldwide to go to market faster, by unlocking the power of quality data in the lab, through chemical digitalisation
We are a big-data and analysis company focused on enabling reproducibility and predictability in chemistry.
We have been perfecting our technology platform over recent years and have been supported and advised by some of the best minds in the industry